Before You Begin (Archived)
To streamline your navigation through our platform, we've structured the content into three main sections: resource storage, resource sharing, and community curation. This arrangement facilitates a logical progression based on your intended actions:
If you're planning to store resources, begin with Before You Begin (Archived) | ResourceStorage
If you already have resources stored and wish to share them, proceed to Before You Begin (Archived) | ResourceSharing
For those interested in reviewing previously curated metadata, proceed to Before You Begin (Archived) | CommunityCuration
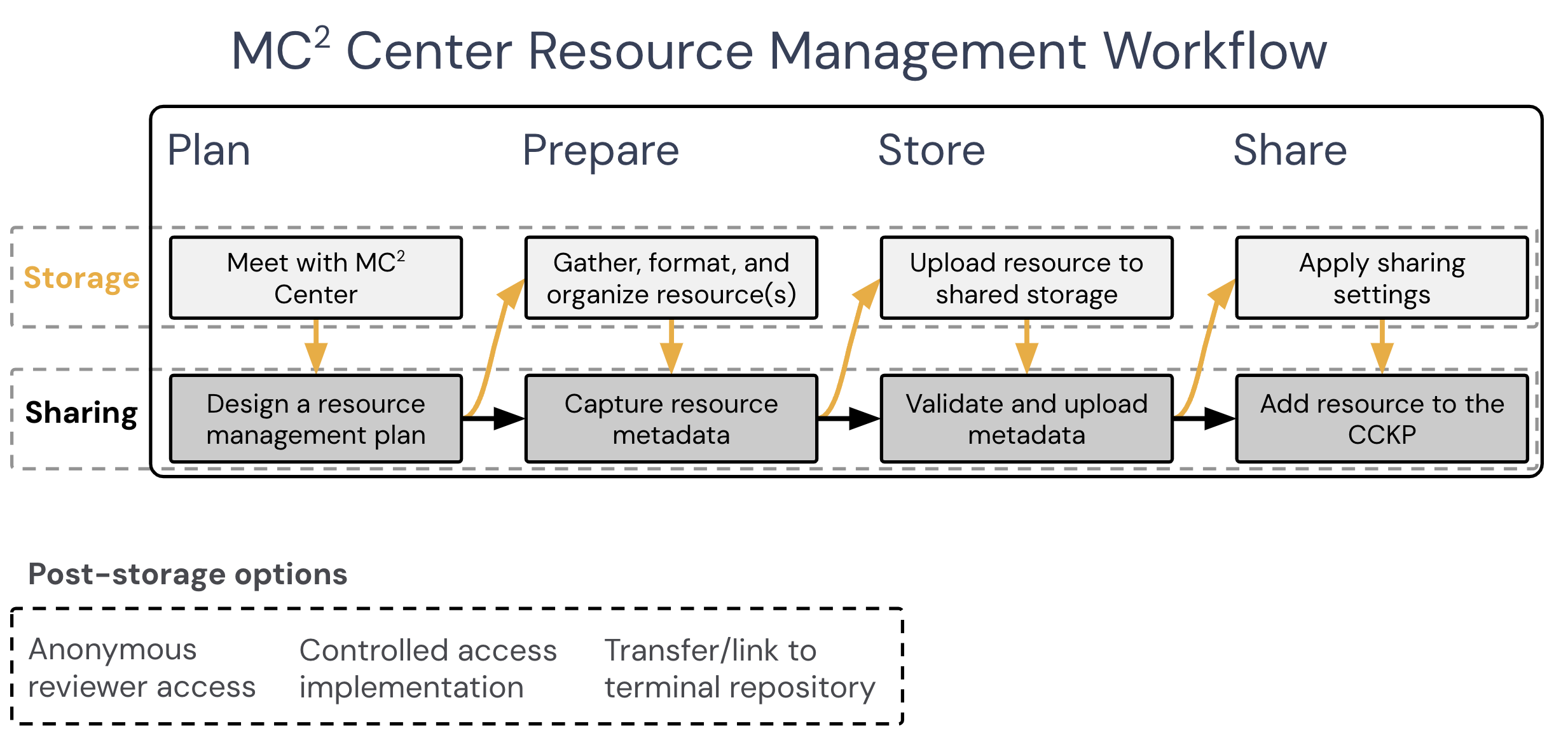
Figure 1. MC2 Center workflow diagrams for Storage and Sharing Paths
Resource Storage
Getting Started: Resource Storage
This page provides instructions for members of the MC2 Center community to begin storing and sharing resources through the MC2 Center.
If you are interested in learning more about the MC2 Center Resource Sharing Workflow, you can access recordings and slides from the introductory webinar here.
If you do not plan to store resources in Synapse, you will be following the Sharing Path of the resource sharing workflow (Figure 1), which is covered in the Resource Sharing section of this page.
1. Synapse Account Setup and Project Access
The MC2 Center has created Synapse Projects for all supported grants to store resources (datasets, computational tools, educational resources, etc.) and associated metadata.
Contributors must have a Certified Synapse profile and must be provided access to their grant’s Synapse Project before uploading or annotating resources.
Please see the complete set of Synapse account setup instructions here: Before You Begin (Archived) | CreateandconfigureyourSynapseaccount
2. Create a resource sharing plan with the MC2 Center
If you intend to store resources in Synapse and you have not previously completed a resource sharing plan for your current study, please schedule a meeting with MC2 Center data manager Orion Banks, via calendly.
More information about resource sharing plans can be found here: Before You Begin (Archived) | Resourcesharingplans
Please email mc2center@sagebase.org if you have any questions or are unable to find a time that works for you.
3. Uploading Resources to your Synapse Project
Once you are a Certified Synapse user and your resource sharing plan has been completed, you are ready to begin uploading resources.
For additional information and guidance on uploading resources, please visit the Resource Management Code section of the Cancer Complexity Knowledge Portal Database in Synapse.
When uploading files to your Synapse project, please do not use the folders listed in Table 1. These folders should only be used to store links to external records or metadata uploaded through the Data Curator Application.
Folders to store your files will be created for you and noted in your resource sharing plan.
4. Annotating Resources with the Data Curator Application
Uploaded resources must be accompanied by descriptions, or metadata, which enable others to understand the data type, content, quality, etc.
The Data Curator Application (DCA) provides an interface to:
Generate spreadsheets (manifests) with columns (attributes) that are defined by metadata models (schemas) contained within the MC2 Center Data Model
Validate annotations (metadata) entered into manifests - validation rules are defined within the MC2 Center Data Model
Store annotations for your resources in your Synapse Project
When trying to identify the appropriate target folder for metadata upload, please refer to your resource sharing plan documents or contact the MC2 Center for support.
Instructions for using the DCA can be found in the MC2 Center Data Ingress Documentation.
You can access the DCA using this link: https://dca.app.sagebionetworks.org/
5. Packaging and release
Whenever possible, resources will be released as open access. Any terms or conditions for accessing the shared resource will be defined in your data sharing plan and implemented prior to release.
The MC2 Center will work with you to organize your shared resource into folders or datasets, which can be listed on the Cancer Complexity Knowledge Portal (CCKP). Please see the resource sharing section of this page for more information on CCKP metadata requirements.
Resource Storage Flow
The MC2 Center is working with investigators to develop recommendations on where to store and how to share the wide variety of data types being generated by this community. Figure 2 provides a brief overview of storing and sharing resources at the MC2 Center.
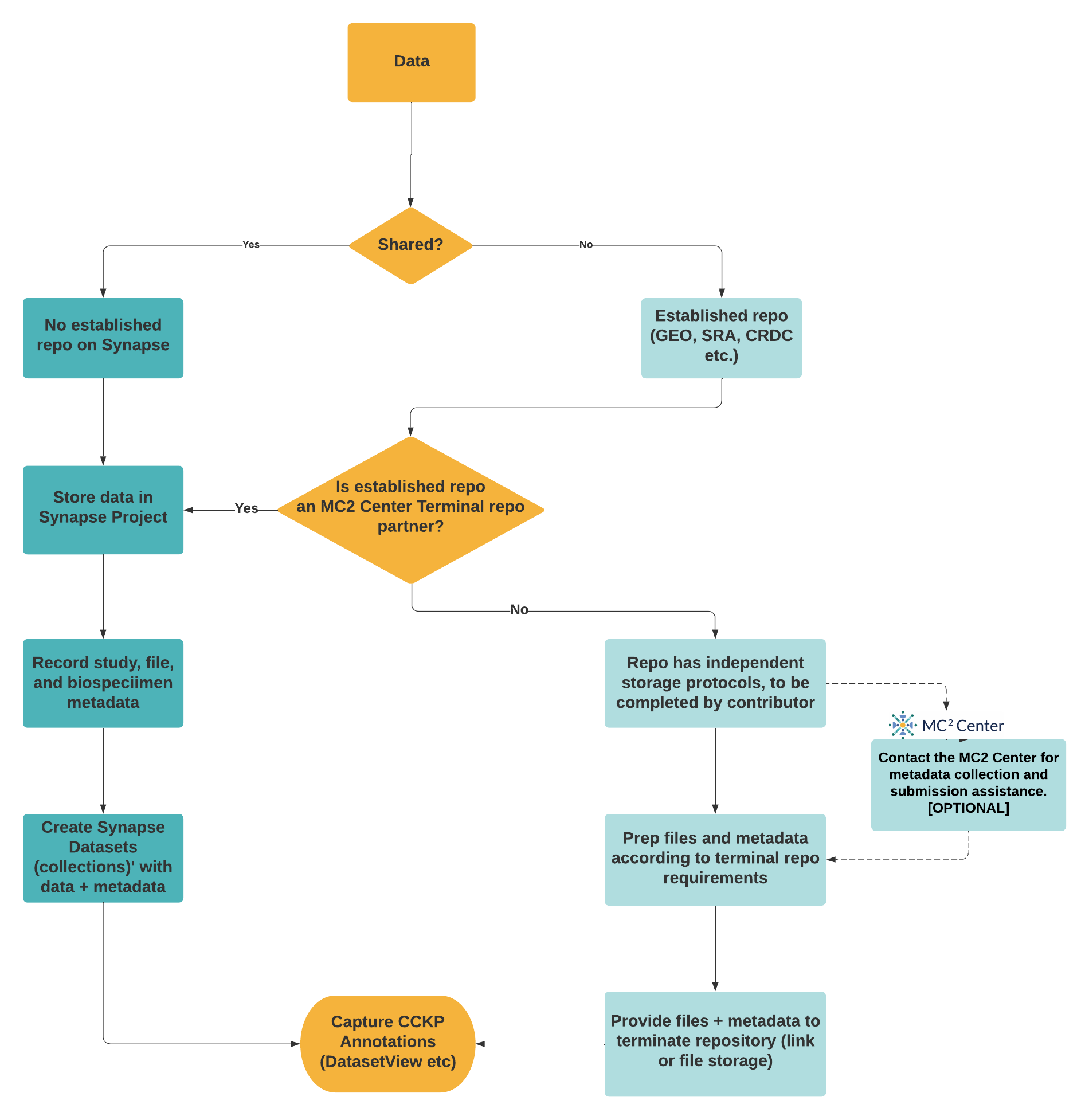
Figure 2. Storing and Sharing Resources through the MC2 Center
Repository Selection
Repositories are typically used for long-term storage of scientific data and other resources. Often, repositories will provide a combination of the following features:
secure storage
permission controls
methods for accessing resources
Whenever possible, the NIH recommends that scientific data is stored in an established repository.
You can explore the sets of NIH-supported Generalist and Domain-specific repositories using the links below.
Generalist vs Domain-specific repositories
Generalist Repositories
Generalist repositories accept data from any scientific discipline. They are designed to be broad in scope and can handle a variety of data types and formats. These repositories are useful for researchers who want to share their data with a wide audience or for those whose data does not fit into a more specialized repository.
Examples:
Dryad: A generalist repository for data underlying scientific and medical publications, Dryad accepts data from any discipline and makes it freely available.
Zenodo: Created by OpenAIRE and CERN, Zenodo is an all-encompassing repository that allows researchers from any field to share datasets, research software, reports, and any other research outputs.
Domain-Specific Repositories
Domain-specific repositories cater to specific scientific disciplines or types of data. These repositories often have specialized tools and metadata standards tailored to the needs of their particular scientific community. They can provide more detailed and appropriate ways to describe and analyze the data, which enhances the usability and discoverability of the datasets within that domain.
Examples:
GenBank: A nucleotide sequence database, GenBank collects and makes available an annotated collection of all publicly available DNA sequences. It’s highly specialized for genetic information.
ClinicalTrials.gov: This repository provides information on publicly and privately supported clinical studies on a wide range of diseases and conditions. It is specifically tailored to the clinical research community.
Example of Use
A researcher working on a new genomic study would likely deposit their data in GenBank to take advantage of the specialized tools and community standards for genetic data. On the other hand, if they have supplementary data that is not genetic (e.g., survey results from participants), they might deposit this data in a generalist repository like Dryad.
By choosing the appropriate repository, researchers can ensure their data is more discoverable and usable by others in their field, or by the broader scientific community.
Resource Sharing
Getting Started: Resource Sharing
This page provides instructions for members of the MC2 Center community to begin sharing resources through the MC2 Center and Cancer Complexity Knowledge Portal (CCKP).
If you are interested in learning more about the MC2 Center Resource Sharing Workflow, you can access recordings and slides from the introductory webinar here.
The content in this section is written for those who have previously stored resources in a publicly accessible repository.
If you intend to store resources in Synapse and have not done so, please return to the previous section of this page, Resource Storage.
1. Synapse Account Setup and Project Access
The MC2 Center has created Synapse Projects for all supported grants to store resources (datasets, computational tools, educational resources, etc.) and associated metadata.
Contributors must have a Certified Synapse profile and must be provided access to their grant’s Synapse Project before uploading or annotating resources.
Please see the complete set of Synapse account setup instructions here: Before You Begin (Archived) | CreateandconfigureyourSynapseaccount
2. Select a relevant metadata template
All individuals supported by an MC2 Center grant are eligible to contribute metadata records to Synapse. Metadata templates for the following resources can be submitted, regardless of which repository they are stored in:
Publication metadata, using the PublicationView template
Dataset metadata, using the DatasetView template
Computational tool metadata, using the ToolView template
Educational Resource metadata, using the Educational Resource template
3. Annotating Resources with the Data Curator Application
The Data Curator Application (DCA) provides an interface to:
Generate spreadsheets (manifests) with columns (attributes) that are defined by metadata models (schemas) contained within the MC2 Center Data Model
Validate annotations (metadata) entered into manifests - validation rules are defined within the MC2 Center Data Model
Store annotations for your resources in your Synapse Project
When trying to identify the appropriate target folder for metadata upload, please refer to Table 1or contact the MC2 Center for support.
Instructions for using the DCA can be found in the MC2 Center Data Ingress Documentation.
You can access the DCA using this link: https://dca.app.sagebionetworks.org/
Why share resources?
Overview
The MC2 Center aims to catalyze collaboration and facilitate an ecosystem of data and resource sharing to create a stable, diverse, and impactful cancer research community.
Figure 3. DCB Consortia supported by MC2 Center
Resource sharing is the process of making your research outputs accessible to others

Figure 4. Potential for data reuse corresponds with the storage method
Resources are the outputs of your research activities and can include:
Experimental data and protocols
Computational tools/code
Publications (pre-prints or peer-reviewed)
Materials for education
Resource sharing benefits
A key benefit of resource sharing is to help accelerate research progress, by reducing the need for others to perform work that has already been done. Additional benefits are noted in Figure 5.
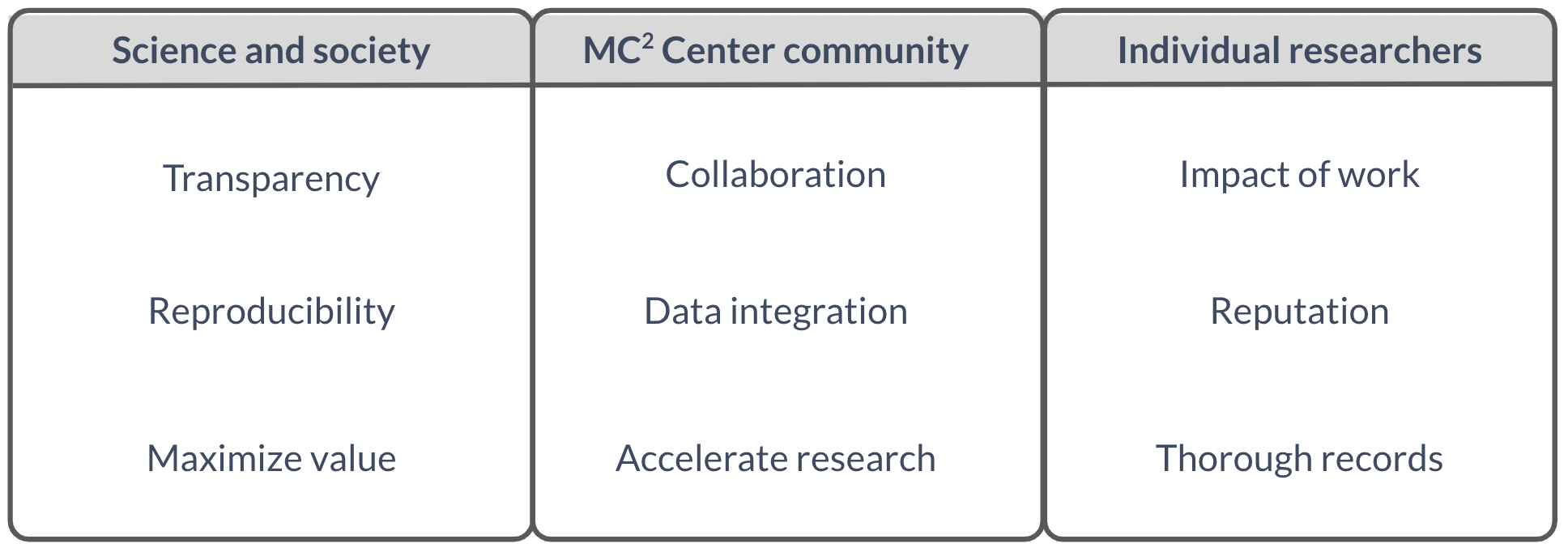
Figure 5. Resource Sharing benefits differ depending on the group under consideration Wilson et al. 2021. FEBS Lett.
Examples of resource sharing
Experimental data sharing has helped create large datasets (e.g., TCGA, INDI) that continuously support research and publication efforts, as shown in Figure 6.
What can you do to encourage reuse of your shared data?
ensure data is findable and accessible
provide thorough descriptions of the data, including how it can be processed, how it was generated, biospecimen information, etc.
ensure your data can be attributed to you, as the original generator
Figure 6. Publications citing data from International Neuroimaging Data-sharing Initiative (INDI), separated by INDI contributors and non-contributors, Milham et al. 2018. Nat Com.
Community Curation
Getting Started: Community Curation
This page is intended to help guide MC2 Center community members through reviewing and contributing metadata to the Cancer Complexity Knowledge Portal database through the Community Curation process.
If you are looking for guidance on storing data or other resources in Synapse, please check out the Resource Storage and Resource Sharing sections of this document.
Who can contribute metadata to the MC2 Center?
All individuals associated with an MC2 Center supported grant are eligible to gain access to a dedicated Synapse project, which is automatically set up for metadata submission.
Once provided access, contributors can add to their grant metadata collection by:
reviewing and updating any existing metadata records
submitting new metadata entries
How to Contribute
Review existing metadata
To help ensure the accuracy and completeness of these resource descriptions, we are asking MC2 Center investigators to review the metadata gathered by the MC2 Center. During review, please correct any errors you notice in the entries and add information to fields that are empty or marked as "Pending Annotation".
Contribute new metadata
All individuals supported by an MC2 Center grant are eligible to contribute metadata records to Synapse. Metadata templates for the following resources can be submitted, as part of the Sharing Only path of the MC2 Center Resource Sharing Workflow depicted in Figure 1:
Publication metadata, using the PublicationView template
Dataset metadata, using the DatasetView template
Computational tool metadata, using the ToolView template
Educational Resource metadata, using the Educational Resource template
Next steps
The following links can be used to help you navigate to the right starting point:
→ I’ve never accessed the Synapse project associated with my grant
→ I’ve previously used the Data Curator Application to upload metadata
Background on MC2 Center resource curation
As part of ongoing efforts to maximize the impact and FAIRness of resources created by NIH/NCI Division of Cancer Biology (DCB) consortia members, descriptions of the publications, datasets, computational tools, and educational materials generated by supported projects are recorded and managed by the MC2 Center. Spreadsheets with predefined sets of columns are used to store the descriptive information, which is commonly labeled as metadata.
The metadata spreadsheets, or manifests, are incorporated into a public Synapse database and made available through the Cancer Complexity Knowledge Portal, which allows users to filter through available resources and identify items that they can repurpose in their work.
The MC2 Center maintains a database of metadata, with information on the publications, datasets, computational tools, and educational resources that are generated by supported grants.
The metadata for each resource is uploaded to grant-specific Synapse projects and subsequently integrated into the Cancer Complexity Knowledge Portal Database. An outline of the MC2 Center resource curation workflow is displayed in Figure 7.
Figure 7. The flow of MC2 Center metadata to the Cancer Complexity Knowledge Portal.
Community Curation Guide
The Cancer Complexity Knowledge Portal surfaces published papers, datasets, tools, and educational resources that MC2 Center curators have annotated.
The initial metadata annotations have been completed by our center and added to the CCKP. For these previously curated resources, we request that you review and provide input on the captured information. Your feedback is highly valuable, and won't require much of your time.
To get started, we encourage you to navigate to the Cancer Complexity Knowledge Portal and confirm the existing resources we have curated and disseminated.
Community curation steps and documentation
Please see below for a complete set of steps and documentation on participating in community curation.
create and configure your Synapse account
access the Data Curator Application
update or record resource metadata
validate and submit metadata with the Data Curator Application
Confirming your resources
https://www.loom.com/share/a7dc474cdf54466fb2d46e6ac241c343?sid=419bbabc-0e4a-4b41-b7f3-631148de5c58Modify your resources
We have different manifests (spreadsheets) for Publications, Datasets and Tools. Please proceed to the respective sections to modify your resources.
For more information visit: Community Curation (Archived)
How-to
Create and configure your Synapse account
These steps can be performed at anytime, but must be completed prior to uploading any files to Synapse, regardless of method (browser interface, command line, or Data Curator App)
Create a Synapse account here
Certify your Synapse account (this step is required to upload data and metadata to Synapse)
Access the Synapse documentation on Certified User Accounts.
Under the “Certified User” section, click on the “Become a certified user” link, which will direct you to a 15-question quiz on the Synapse Commons Data Use Procedure.
Request access to your grant’s Synapse Team
Access this public Synapse Table, containing information about MC2 Center grants
Identify the row matching your grant and scroll to the far right columns
Use facet filters or the table search function to limit the number of entries in the table. Please see Synapse documentation on searching tables for more information.
Access the Synapse page for your team, using the link in column “grantSynapseTeam”. You can learn more about Synapse Teams here.
Click on the “Request to Join Team” button.
Once added to the Team, you will have edit-level access to your grant’s Synapse project.
Access the Synapse project for your grant
Follow steps 3a - 3c. For 3c, your Synapse project can be accessed via the link in column “grantSynapseProject”
Once your join team request has been approved, you should have access to the full functionality of your Synapse project.
Selecting a folder for metadata upload
Please use the table below to identify the metadata model and folder name associated with each type of resource curated for the Cancer Complexity Knowledge Portal. If the resource you would like to annotate is not listed below, please refer to your resource sharing plan documents or contact the MC2 Center for support.
When uploading resource files to your Synapse project, please do not use the folders listed in Table 1. These folders should only be used to store links to external records or metadata uploaded through the Data Curator Application.
Folders to store your files will be created for you and noted in your resource sharing plan.
Table 1. Metadata template and storage folder reference for MC2 Center contributors
Resource type | Metadata template name | Target folder name in Synapse project | Metadata template details |
Publication | PublicationView | publications | |
Dataset | DatasetView | datasets | |
Computational tool | ToolView | tools | |
Educational resource | EducationalResource | education | Education Resource Data Model - MC2 Center Data Models Explorer |
Resource sharing plans
The MC2 Center uses template-based documents (resource sharing plans) to record information about your project, Synapse access details, and files you plan to store in Synapse. These documents will be shared with you during your intake meeting. Links to form templates can be found in Table 2, below.
MC2 Center resource sharing plan documents should be used as reference materials, as they contain critical information to help you annotate your resources, upload your files, and submit metadata to the appropriate folders in your Synapse project.
During your intake meeting, if time allows, you will work with an MC2 Center representative to add information about the datasets, computational tools, and educational resources you plan to store in your Synapse project.
If you have previously created a resource sharing plan and are unable to access the document(s), please contact the MC2 Center at mc2center@sagebase.org.
Table 2. Directory of MC2 Center Resource Sharing Plan form templates
Form type | Blank template link |
Intake | |
Dataset sharing plan | |
Computational tool sharing plan | |
Educational resource sharing plan | [Template] MC2 Center Resource Sharing Plan - Educational Resources |
